Cotton genome sequencing is changing the way cotton is grown, helping improve fiber quality, pest resistance, and drought tolerance. By mapping the complete DNA of cotton species, researchers can identify genes tied to better traits and use them to create high-performing varieties for U.S. farmers. This means cotton plants can be tailored to thrive in specific climates, reduce pesticide use, and increase yields.
Key takeaways:
- Modern cotton varieties now produce finer fibers and higher yields (up to 1,225 pounds per acre).
- Advanced sequencing tools, like PacBio and CRISPR, allow precise genetic improvements.
- Researchers have identified over 25,000 new genes, including those for fiber strength, drought resistance, and pest control.
- New breeding techniques have drastically shortened the time needed to develop improved cotton varieties.
These advancements ensure cotton farming stays productive and resilient, even as climate and agricultural challenges grow.
New Technologies in Cotton Genome Sequencing
Next-Generation Sequencing Methods
The transition from older sequencing technologies to long-read platforms has completely reshaped cotton genome research. Tools like PacBio CLR and HiFi now provide the depth needed to analyze cotton's genome, which is 70.8% repetitive. Notably, Ty3 repeats account for 34.3% of this genome, covering 776.5 million base pairs.
In May 2024, a team led by Avinash Sreedasyam utilized 116.7× PacBio CLR coverage combined with 172× Hi-C scaffolding to create chromosome-scale reference genomes for three modern cultivars: 'UGA230', 'UA48', and 'CSX8308'. This method corrected 35 chromosomal inversions, spanning 122 million base pairs, that had been misassembled in earlier studies. Meanwhile, Oxford Nanopore Technologies has advanced telomere-to-telomere (T2T) assemblies, producing seamless views of complex genomic regions like centromeres. In August 2024, researchers Gai Huang and Yuxian Zhu achieved a near-complete T2T genome for Gossypium raimondii, identifying 13 centromeres and telomeres on 25 chromosomal ends.
"The corrected inversions, increased per-base sequencing depth, improved accuracy and substantial reduction in gapped sequence in the v3 TM-1 genome result in a superior reference genome that will better support breeding and biotechnology goals." – Nature Plants
These advancements lay the groundwork for linking genomic variations to specific cotton traits with greater precision.
How Advanced Sequencing Techniques Are Used
With these improved assemblies, researchers are now using sequencing methods to directly connect genetic variations to important traits. By assembling genomes and identifying variations, scientists can uncover genetic factors that influence key characteristics. Graph-based pan-genomes, for instance, capture structural variations (SVs) and presence-absence variations (PAVs) across hundreds of cotton accessions. This approach reveals genetic diversity that single-reference methods often miss. A study involving 107 genome assemblies identified specific loci tied to disease resistance that earlier SNP-based methods failed to detect.
Functional Haplotype (FH) mapping takes this a step further by organizing variants into gene-level units, minimizing interference from linkage disequilibrium in cotton's polyploid genome. In early 2025, researchers applied FH mapping to 3,724 cotton accessions, identifying 532 quantitative trait genes (QTGs) with strong breeding potential. They even validated a fiber quality QTG, which encodes ferulic acid 5-hydroxylase 1, using CRISPR-Cas9 experiments. This level of precision allows breeders to target traits like fiber strength, drought tolerance, and pest resistance, helping to develop superior cotton varieties tailored for U.S. agricultural needs.
sbb-itb-0e617ca
Utilizing Targeted SNP-Based Genotyping to Rapidly Introgress Interspecific Germplasm
Genetic Traits Identified Through Sequencing
Recent advancements in sequencing technologies have made it possible to link specific genetic traits to key agricultural improvements in cotton production.
Pest Resistance
Sequencing has been instrumental in identifying genes that enhance cotton's resistance to pests, cutting down the need for pesticides. For instance, the GhnsLTPsA10 gene has been shown to combine resistance to both diseases and insects. In the fight against whiteflies, researchers used Genotyping-by-Sequencing to locate a major quantitative trait locus on chromosome A04 (qWC1_A04) and identified two key genes, At4g27190 and RPPL1, that are responsible for resistance. These breakthroughs enable marker-assisted selection, allowing breeders to identify pest-resistant plants without relying heavily on field trials.
In October 2018, a collaboration among the University of Arizona, University of Tennessee, and Nanjing Agricultural University made a groundbreaking discovery about pest adaptation. The team, led by Professors Bruce Tabashnik and Yidong Wu, pinpointed a single DNA base pair change in the HaTSPAN1 gene of the cotton bollworm (Helicoverpa armigera) that provides dominant resistance to Bt cotton. Their analysis of thousands of moths collected over a decade (2006–2016) revealed that the mutation's frequency skyrocketed from 1 in 1,000 to 1 in 10. This prompted Chinese farmers to adopt cotton varieties producing multiple Bt proteins to maintain pest control effectiveness.
"Without the latest advances in genetic technology, it would not have been possible to find the single DNA base pair change causing resistance among the hundreds of millions of base pairs in the bollworm's genome." – Bruce Tabashnik, Regents' Professor, University of Arizona
Drought Tolerance
Drought resistance has become a major focus for researchers, especially as unpredictable rainfall increasingly impacts U.S. cotton growers. Sequencing has revealed genetic markers that help cotton plants thrive in water-scarce conditions. Drought stress has been shown to reduce key growth metrics like shoot length (61%), root length (52%), and photosynthetic rates (60%). High-quality reference genomes for modern cultivars such as UGA230 and UA48 are now being used to breed varieties suited to specific regional climates, from the southeastern Cotton Belt to higher-latitude fields.
Some genotypes, including IR-14 and FH-540, have demonstrated remarkable drought tolerance, maintaining superior photosynthetic efficiency under stress. These varieties also show a 2.3-fold increase in superoxide dismutase activity and a 57% increase in proline content during severe drought conditions. Additionally, sequencing of G. barbadense (Pima cotton) has identified genomic regions that control photoperiod sensitivity, enabling breeders to incorporate tropical germplasm into U.S. breeding programs. Pima cotton, though only 3% of U.S. production at 426,000 bales annually, commands higher prices due to its superior fiber quality, which remains stable even under drought stress in regions like the Western High Plains.
Improved Fiber Quality
Genome sequencing has also unlocked hundreds of genes tied to fiber characteristics, boosting both profitability for farmers and quality for textile manufacturers. In January 2026, researchers Li Fuguang and Yang Zhao'en from the Institute of Cotton Research of the Chinese Academy of Agricultural Sciences published a "super pangenome" study in Nature Genetics. The study analyzed 107 upland cotton varieties and identified 69 genetic loci linked to fiber quality and yield, including 62 previously unknown markers. They also pinpointed the VWD11 region, which provides resistance to Verticillium wilt. Structural variations, such as inversions on chromosomes A06 and A08, have been directly associated with improvements in fiber length and strength.
Modern cultivars like UA48 and UGA230 now produce much finer fibers, with a mean circumference of about 53.23 μM, compared to the historical TM-1 reference of 70.8 μM. These cultivars also deliver higher lint yields, ranging from 950 to 1,225 pounds per acre, far surpassing the older standard of 737.83 pounds per acre. These advancements not only enhance fiber quality but also improve yield efficiency, giving U.S. cotton farmers a stronger position in the global market.
New Breeding Techniques Enabled by Genome Research
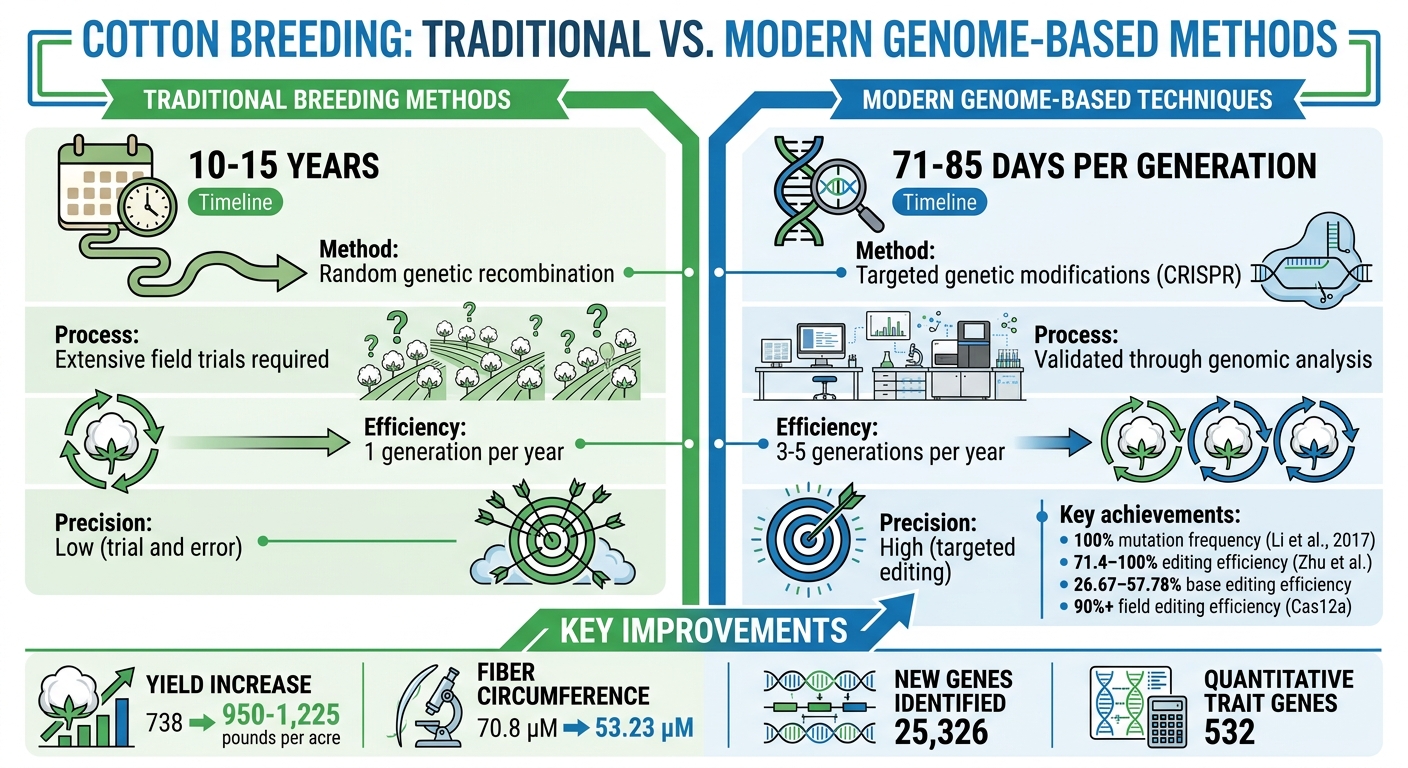
Traditional vs Modern Cotton Breeding: Timeline and Efficiency Comparison
Genome sequencing has turned cotton breeding into a precise and efficient science. While traditional breeding methods often took 10–15 years and relied on random genetic recombination and extensive field trials, modern genome-based techniques have drastically shortened this timeline. Researchers can now make targeted genetic modifications and validate results much faster, transforming how new cotton varieties are developed for American farms.
CRISPR and Genome Editing
CRISPR technology has become a game-changer in cotton improvement, allowing scientists to modify specific genes with a level of precision that was previously unimaginable. The cotton genome, with its 2.5 billion base pairs and extensive gene duplication, presents unique challenges. However, high-quality reference genomes now guide researchers in designing single guide RNAs (sgRNAs) that target precise genes without causing unintended changes.
"Genome editing technology, specifically with clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein (Cas) tools, has opened new possibilities for trait development in cotton. It allows precise and efficient manipulation within the cotton genome when compared with other genetic engineering tools." – Journal of Cotton Research
The potential of CRISPR is already evident. In 2017, Li et al. achieved a 100% mutation frequency in cotton by using CRISPR/Cas9 to knock out the GhMYB25-like genes without off-target effects. Similarly, Chen et al. used the technology to modify the GhFAD2 gene family, leading to cottonseed oil with higher oleic acid and lower linoleic acid levels - resulting in a healthier, more stable oil free of foreign DNA. Zhu et al. targeted the GhAlaRP gene, achieving mutation efficiencies of 71.4% to 100% in the A subgenome and 92.9% to 100% in the D subgenome, advancing research into fiber development.
Recent advancements in CRISPR systems include base editing, which allows single DNA changes without cutting the strand. This method has achieved efficiencies between 26.67% and 57.78% at specific genome locations. Cas12a systems have shown over 90% editing efficiency under field conditions. In addition, multiplex editing enables the modification of multiple genes at once. When paired with "speed breeding" techniques - using controlled environments and extended light periods - generation times can drop to 71–85 days, allowing for 3–5 generations per year instead of the typical single cycle.
"When haplotype analysis is combined with genome editing, breeders are no longer just passively screening but can actively design. They can directly superimpose beneficial genes on superior germplasm to create new varieties." – Zhen Li, Hainan Institute of Biotechnology
These advancements provide a foundation for integrating broader genetic diversity into breeding programs.
Mutant Libraries and Screening Tools
In addition to CRISPR, mutant libraries created through chemical and physical mutagenesis offer another way to uncover beneficial genetic variations. These libraries are especially useful for recovering traits lost during cotton's domestication, which reduced genetic diversity to less than 10% of what is found in wild relatives. Ethyl Methane Sulfonate (EMS) is used to create point mutations that can be incorporated into elite lines, while electron beam irradiation has generated unique traits like novel lint colors and ultrafine fibers.
Genome sequencing has further streamlined this process with "QTL-seq" pipelines. By sequencing pooled DNA from plants with desirable traits and comparing it to reference genomes, researchers can pinpoint the genetic changes responsible for improvements. This method has been used to increase fiber length by up to 12%.
To validate these new traits, high-throughput phenotyping tools come into play. UAV-based drones collect aerial images to monitor plant health, growth, and stress responses across large fields. Advanced Fiber Information Systems (AFIS) provide precise measurements of fiber quality, though their throughput is lower than traditional High Volume Instrument (HVI) systems, which offer faster but less detailed assessments. Together, these tools enable the creation of "digital twins" - virtual models that combine genomic, phenotypic, and environmental data to predict crop performance. This reduces the risks and costs associated with bringing new cotton varieties to market.
Applications for U.S. Cotton Farming
Advances in cotton genomics are bringing tangible benefits to American cotton farmers, transforming how they choose crop varieties, manage resources, and adapt to diverse regional climates.
Breeding for U.S. Climate Conditions
With the help of genomic advancements, breeders in the U.S. are now developing cotton varieties specifically tailored to local climate demands. A joint effort between U.S. and Australian breeding programs has resulted in elite cultivar genomes that outperform the long-standing TM-1 reference. For example, "UA48" was crafted for areas with shorter growing seasons, offering early maturity and robust resistance to bacterial blight - key traits for regions with variable climates. On the other hand, "UGA230" is designed for the Southeastern U.S., where longer growing seasons allow it to produce yields ranging from 950 to 1,225 pounds per acre.
These breakthroughs have uncovered 25,326 previously unknown genes, giving breeders precise targets for improving traits like disease resistance and climate adaptation. As highlighted in a study:
"TM-1 has served the cotton community well as the reference genotype since 1970 but is no longer used in any breeding programmes because of its inferior yield and fibre quality traits compared with modern germplasm and cultivars." – Nature Plants
Resource Efficiency Through Genetic Improvements
Genetic advancements are also driving more efficient use of resources, a critical need given the rising costs of water, fertilizer, and pesticides. For instance, Pima cotton (G. barbadense) shows natural drought tolerance, maintaining fiber quality even under water-scarce conditions in areas like the Western High Plains. By incorporating resilience genes, irrigation requirements can be reduced without compromising yield. Additionally, disease resistance genes have a direct impact on resource efficiency. Both "UA48" and "CSX8308" demonstrate strong resistance to bacterial blight, while "CSX8308" also resists Fusarium wilt. Early-maturing varieties like "UA48" further lower water and fertilizer needs by shortening the growing season.
A specific genetic breakthrough involves the VOZ1 gene at the Gb_Ppd locus, which overcomes photoperiod sensitivity. This allows breeders to incorporate tropical alleles that improve nutrient and water efficiency.
Research Partnerships Supporting Cotton Growers
To ensure these advancements reach the fields, research partnerships are playing a vital role in connecting laboratory discoveries with practical farming applications. The International Cotton Genome Initiative (ICGI), supported by Cotton Incorporated, acts as a key platform for translating genomic research into breeding innovations. At the May 2018 ICGI conference in Edinburgh, Scotland, Dr. Don Jones of Cotton Incorporated led discussions on applied genomics, calling the event a pivotal moment for integrating genomic progress into agricultural practices.
The USDA Agricultural Research Service (ARS) also contributes significantly through national programs aimed at developing region-specific cultivars. Academic institutions like Iowa State University are further advancing the field by conducting large-scale genomic studies, including the exploration of structural variations using graph pan-genomes, which go beyond traditional SNP analysis.
Conclusion
Cotton genome sequencing has reshaped how American farmers enhance their crops. What was once a slow and uncertain process has become a precise, data-driven approach. By replacing the outdated TM-1 genome with modern cultivar genomes, researchers have uncovered over 25,000 genes, driving yields up from approximately 738 pounds per acre to an impressive 950–1,225 pounds per acre. These advancements have led to greater profitability, thanks to higher yields, lower input costs, and crops better suited to regional climate challenges.
The identification of 532 quantitative trait genes and the use of genomic selection have made it possible to address long-standing challenges, such as balancing fiber length and strength, improving drought tolerance, and reducing pesticide use. For example, in 2025, researchers demonstrated that introducing a single favorable allele of GhROPGEF5 increased fiber length by about 1.5 mm and improved fiber strength by roughly 1.8 g tex⁻¹ - without compromising yield. Achievements like this highlight the precision and efficiency of modern genomic techniques, which far surpass traditional breeding methods. These results emphasize the importance of ongoing collaboration across institutions.
Organizations like the USDA Agricultural Research Service, Cotton Incorporated's International Cotton Genome Initiative, and universities play a crucial role in advancing this research. Their efforts in exploring wild germplasm, validating new genes with tools like CRISPR, and bringing lab discoveries into the field are essential to the future of cotton farming.
As climate challenges grow more pressing, the need for innovation has never been clearer. Events like the devastating 2011 Texas drought, which wiped out more than half of the state's cotton crop and caused significant financial losses, serve as a stark reminder of the stakes. Genomic research is no longer optional; it is the backbone of a resilient and competitive cotton industry. By investing in cutting-edge sequencing, precise genomic references, and strong partnerships, the U.S. cotton sector can remain competitive while embedding sustainability into every acre planted.
FAQs
How soon will genome-based cotton varieties reach U.S. farms?
Genome-based cotton varieties are anticipated to make their way to U.S. farms within the next few years. Research from early and mid-2026 has showcased progress in cotton genomics, which will play a crucial role in shaping breeding strategies and developing new cultivars. These advancements target major farming challenges, including improving pest resistance, enhancing drought tolerance, and boosting fiber quality. The result? A step toward more efficient and environmentally friendly cotton production.
Will CRISPR-edited cotton be regulated or labeled differently?
CRISPR-edited cotton in the U.S. is likely to face regulation and may need specific labeling under current guidelines. U.S. regulations, along with those in Europe, require oversight and labeling for CRISPR technology to ensure it aligns with agricultural standards.
How can growers choose varieties matched to local drought and pest pressure?
Growers now have the tools to choose cotton varieties tailored to their specific environments, thanks to genomic research that highlights traits such as drought tolerance and pest resistance. Breakthroughs in cotton genome sequencing have pinpointed the genes responsible for these traits, enabling breeders to create cultivars designed for particular regions. By applying this knowledge, growers can select varieties that are better suited to their local climate and pest pressures, leading to improved yields, higher fiber quality, and more efficient farming practices.
Related Blog Posts
- How CRISPR Gene Editing is Engineering the Future of Drought-Proof Cotton Yields
- Latest GM Cotton Seed Genetics Research: What’s Reshaping Global Supply
- CRISPR Technology in Cotton Seed Genetics: Latest Research and Farmer Applications
- Genetically Modified Cotton Research: Innovations Driving Higher Yields and Pest Resistance


